Are Covid-19 PCR tests detecting human and microbial genetic material?
All genetic sequences used in the widely-used CDC PCR test are commonly found in human and microbial DNA, which is likely generating significant numbers of false positive test results; the degree to which false positives are being generated due to this serious design flaw is an empirical matter that requires additional research
Dr. Sin Lee , an expert in PCR testing and genetic sequencing, reviewed this essay and agrees with its contents and conclusions.
The Washington Post published in late December 2020 a detailed and alarming retrospective look at how the US CDC developed its new PCR test for detecting the 2019 novel coronavirus, now known as SARS-CoV-2 (Vice News published on the same story far earlier in 2020).
We learned that CDC decided to develop its own new test with three “probes” (genetic sequences thought to be unique to the new virus) in order to have an accurate test that avoided false negatives and false positives. Other tests often used only one or two probes but CDC wanted to use a better test with three probes; as the Post article states: “the agency was seeking a more sophisticated test.”
However, even though the FDA approved the new CDC test Feb. 4 2020 under its Emergency Use Authorization (EUA), CDC didn’t send the final test to labs until the very end of February. This is because during troubleshooting in various labs in February, the new test kept producing high numbers of false positives in lab samples that were known to not contain the novel virus. It wasn’t clear at the time what was causing these false positives.
ProPublica published an investigation into CDC’s highly problematic test development process, finding that:
Complaints poured in as soon as the tests arrived at the public health labs. Before screening any samples from patients, scientists checked to ensure the tests worked, using water for a negative and the stand-in for the virus for a positive. They found the same problem with the third target: It registered as positive when just testing water.
“There is likely a widespread issue that will need to be addressed immediately,” a California public health official said in an email to the CDC on Feb. 8.
The same story includes a big hint as to what probably happened (emphasis added):
Lindstrom [CDC’s test development leader] told colleagues he was convinced there was contamination, but some CDC leaders insisted that the problem was actually a faulty design akin to a software bug — that Lindstrom had chosen genetic sequences that could cause a glitch and show a false positive, according to emails and interviews.
CDC decided to drop the third probe, which was thought to be the likely cause of many of the high numbers of false positives, and ordered all new versions of the test. So rather than use all three probes, labs in the US have been using two probes thought to be unique to the novel virus for much of 2020 (both probes are part of the virus’s N gene that codes for nucleocapsids, a kind of shell that attaches to the RNA of the virus).
The Vice story linked above says much the same thing, quoting Tomer Altman, a Stanford-trained bioinformatics specialist:
“I just put the primers as input into this program and within seconds it tells you there’s problems with it,” Altman said. “It’s freely available. It’s not hard to use at all. It’s one of those things you’d do as a basic quality control check. It seems like they hand-designed these primers and it seems like they didn’t use computer tools to check it.”
CDC’s website hints at similar information here but omits mention of the large numbers of false positives that were found during troubleshooting of the new test in February of 2020, and doesn’t mention that false positives may still be likely (accessed Jan. 8, 2021):
Before laboratories use a new test on samples from patients, they must verify the test performance (make sure it works as expected) using “positive” and “negative” control materials. The positive control should always test positive, and the negative control should always test negative. During validation of the CDC SARS-CoV-2 test, some laboratories discovered a problem with one of the test’s three reagents — chemicals required to run a test. The reagent produced a positive result with the negative control, so laboratories could not verify test performance.
To resolve the issue, CDC laboratories determined that this reagent could be left out without affecting test accuracy because of the built-in redundancy in test design. The redundant design saved time by allowing the kits to be used without the reagent. FDA authorized this modification, and new test kits with the two necessary reagents were manufactured and distributed to states. These kits are still in use.
This fix was thought to render the test sufficiently accurate to detect the virus without too many false negatives or false positives.
All three of CDC’s original test probes were parts of the virus’s N gene (the probes were labeled N1, N2 and N3). The test that ended up being used widely in the US uses only the N1 and N2 probes, plus a human DNA probe known as RNase P, which I’ll discuss below.
Once the test was rolled out nation-wide in early March we quickly began to see Covid-19 case counts rise dramatically. In fact, testing and case counts correlate fairly well for the first two months, as Figure 1 shows. This is to be expected because the CDC “case definition” defined a “confirmed” case of Covid-19 simply as a positive lab test (almost all previous respiratory ailment case definitions have required symptoms plus a lab test for a confirmed case and it’s not clear why CDC opted for this far looser definition of a confirmed case for Covid-19).

Did CDC roll out a fundamentally flawed PCR test?
But was the revised test accurate? Were the positive test results actually finding cases of Covid-19 or were they picking up in most cases genetic material that exists widely in the human genome and in microbes that commonly live in humans?
Is this why there were so many false positives in testing the CDC’s PCR test in February? As Willman reported for the Washington Post it wasn’t just the N3 probe (which was ultimately dropped from the final test) but also the N1 probe that was yielding a lot of false positives during lab troubleshooting.
The National Center for Biotechnology Information (NCBI) maintains an impressive database of genetic information from numerous species, including humans and thousands of microbes and viruses. It’s public and can be searched with a relatively simple interface called BLAST.
I used the BLAST search function to look for all of the primers and probes used in the CDC test, which is a total of 9 sequences in the test as approved (N1, N2 and RNase P).
What I found should be surprising: All primers and probes are commonly found in human DNA and microbial genetic material, in sequence lengths far longer than the 10 bases required for a probe to bind. Even though probes and primers are far longer than 10 bases in length it is sufficient (Ryu et al. 2000; Garhan et al. 2013) for a match that only 10 contiguous bases are an exact match, or even as little as six for primers at certain temperatures — these key pieces of information were confirmed in a dialogue with Dr. Sin Lee, a pathologist who operates a DNA sequencing lab in Connecticut. I discuss his work further below.
All CDC PCR test primers and probes are common in human DNA and microbes
The following table shows all of the CDC PCR test primers and probes, omitting the N3 probe for reasons discussed above. The site includes a disclaimer that “These sequences are intended to be used for the purposes of respiratory virus surveillance and research.” They “may not be used for viral testing under FDA’s authorization of the CDC 2019-nCoV Real-Time RT-PCR Diagnostic Panel.”

I’ll provide some examples of these matches, without being exhaustive.
Figure 3 shows the microbe search results for the N1 gene forward primer (20 bases long: gaccccaaaatcagcgaaat) from the CDC PCR test, with a 19-base perfect match in the top example and an 18-base perfect match in the second example.
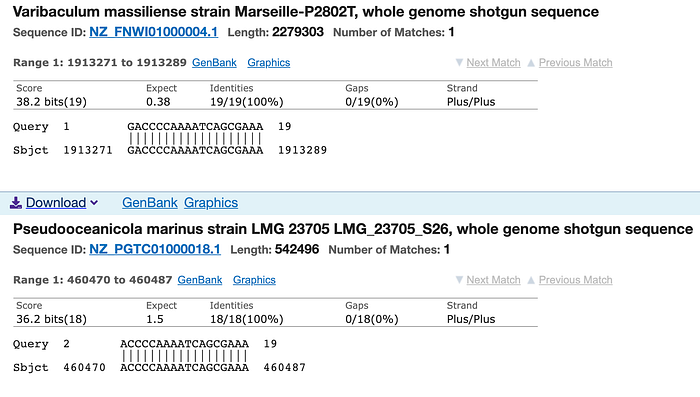
FDA’s May 2020 guidance for testing for Covid-19 states that any homology over 80% (meaning more than 80% of the sequence matches) is problematic in terms of cross-reactivity “on common respiratory flora and other viral pathogens.” Finding 19 out of 20 bases matched for this N1 forward primer is far higher than 80% homologous and is, accordingly, highly likely to result in cross-reactivity.
Figure 4 shows the human DNA match for the 28-base N1 gene probe (FAM-ACC CCG CAT TAC GTT TGG TGG ACC-BHQ1), with a maximum of 15 bases perfectly matching in human chromosome 2. This is not 80% homologous but is still far more than the 10–12 base match required.
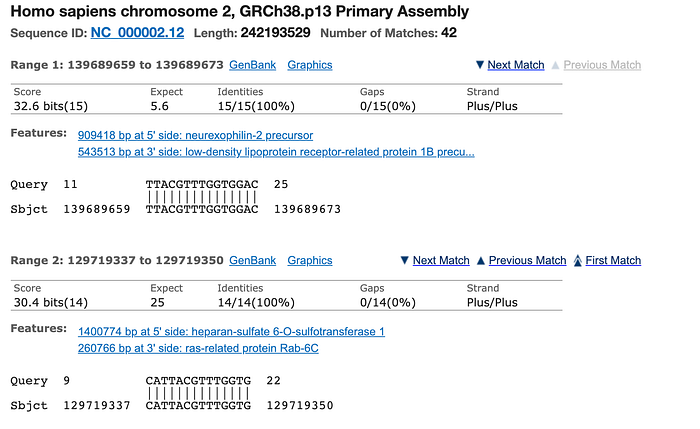
The CDC PCR test 27-base N2 probe yields 97 human DNA matches up to 21 out of 22 perfect matches and 15/15 perfect matches at the high end of the list of matches.

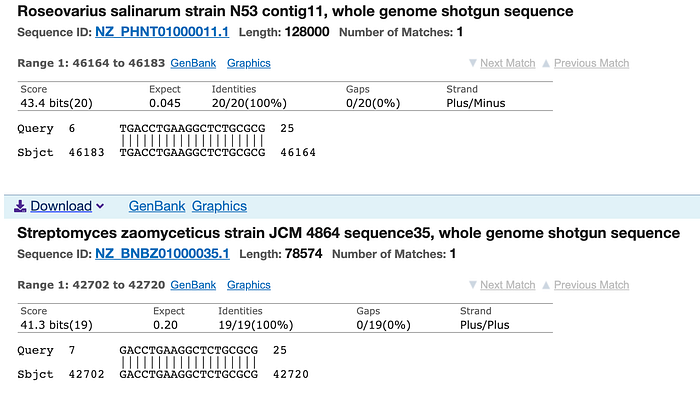
The same N2 probe yields hundreds of microbial matches, up to 19 out of 19 perfect matches.
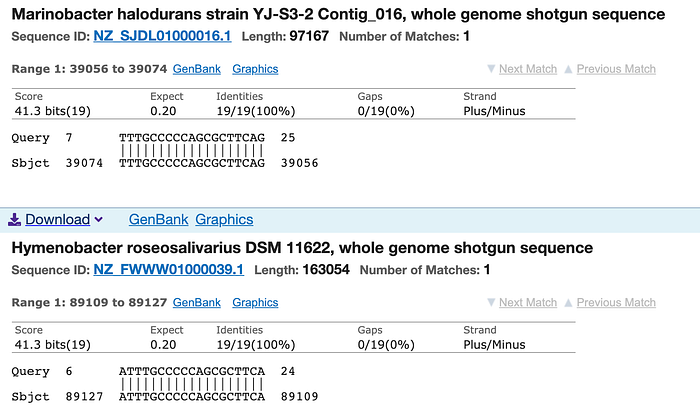

Last, the third probe, RNase P, was included as a way to demonstrate that samples were in fact coming from humans, as a way to confirm that the tests were being used on human samples. However, it turns out that the RNase P probe is not only, as expected, present in human DNA (of course, since that’s how it was designed; my search found a perfect 23/23 match in chromosome 10) but also relatively common in microbes, with up to 20 out of 20 perfect matches for this 27-base sequence, and dozens of matches for sequences 10 or more bases long.
What does all of this mean and what do we do about it?
As mentioned above, matching only 10 bases in the probe genetic material is enough to trigger amplification and a false positive. I’ve demonstrated that all of the probes show far more than 10 bases matching in hundreds of human chromosome sequences and microbes.
Matching with just one of the probes was not enough in most cases to trigger a positive result, because all three probes are supposed to be positive for a positive result for the CDC PCR test (see FDA instructions for use for the CDC PCR test here (p. 37)). However, other tests developed after CDC’s test require only one probe match to be considered a positive result. The Cepheid test, for example, states that only one probe is required to be a “presumptive positive.” This contradicts the point of CDC’s “smart test,” but thus far it seems that there has been little to no outcry about the potential for false positives with these later-developed single probe tests.
Also, we have data demonstrating that many labs in the UK have been requiring only one positive probe result for a positive test (as high as 38% of all UK positive PCR test results early in 2021), so if even one of the probe results is a false positive it will in these cases yield a false positive test result. It is not clear at this point if any US labs are doing the same thing as these UK labs in requiring only one N gene positive for an overall positive test result, but it is reasonable to expect that they may be since the US and UK share very similar public health surveillance and testing systems.
A recent development, in April of 2021, validates this concern. FDA issued guidance that a number of PCR tests may be less effective at detecting new variants like B.1.1.7 and as a result FDA advises that only one probe positive may be sufficient for a presumptive positive test result for a number of tests listed on FDA’s website, including the commonly-used Cepheid test just mentioned.
So how many of the positive test results in the US are in fact false positives due to the pre-existing human and microbial genetic material? The bottomline is that we don’t know yet because to arrive at a firm answer we’d need to perform genetic sequencing (Sanger sequencing) on positive test results conducted so far, or at least a good sample thereof in order to arrive at a reasonable estimate.
Dr. Sin Lee, mentioned above, has written extensively on these issues, and in dialogue with him about my findings with the BLAST database he opined that 70–90% of the positive PCR test results could indeed be false positives resulting from this preexisting genetic material. He told me that “A good PCR positive test may be >90% correct, in my hands. But the RT-qPCR positives are probably 70–90% false positives in practice.”
His proposed solution, described in a pair of peer-reviewed publications (here and here), is to use Sanger sequencing to test for the SARS-Cov2 virus, looking for a 398-base sequence that Lee suggests is mostly conserved even with the growing number of variants of the virus. He told me that “the [genetic] sequence must be long enough to be unique beyond a reasonable doubt. We do not know which RT-qPCR sample in fact contains SARS-CoV-2 genetic materials.”
The general view is that Sanger sequencing is too expensive and time-consuming to substitute for PCR testing, but Dr. Lee feels that it could be scaled up to be able to fix the fundamental issues facing the PCR tests. The cost currently is around $150 if done at local hospitals — about the same as the actual cost for PCR tests — and as Sanger sequencing becomes more widespread that cost will come down.
And as concern about virus variants grows, Sanger sequencing is becoming a more common part of the surveillance testing system in the US and other countries (the Biden administration and Congress have approved $billions in new funding for sequencing of test results in order to track variants). This could be a good dynamic in terms of Sanger sequencing becoming a key part of the confirmation process for PCR tests more generally.
Can the presence of PCR test primers and probes in human DNA and microbes explain some outstanding mysteries?
In sum, Dr. Lee and others have demonstrated that the PCR tests are fundamentally flawed and leading to highly misleading results. The degree to which false positives are being generated may go far in explaining the various anomalies regarding testing positive one day and then negative the next, and the degree to which some labs have been found to give up to 100% false positives.
For example, a high profile incident involved 77 NFL players testing positive for Covid-19 in a New Jersey lab, only to all be negative upon re-testing. NFL.com reported in August of 2020:
Testing irregularities at one of the labs used by the NFL led to 77 positives for COVID-19 among players and staff members from multiple teams Saturday. NFL Network Insider Ian Rapoport and NFL Network’s Tom Pelissero reported that all 77 original tests were rerun Sunday night, and every single one came back negative, per sources informed of the situation.
Another series of anomalous tests came from research labs on the East Coast. The New York Times reported in a November 2020 story, “These Researchers Tested Positive. But the Virus Wasn’t the Cause,” that a series of positive PCR test results seemed to result from environmental RNA rather than actual infections. The Times reported:
“It was everywhere,” said Gabriel Filsinger, a member of the Church lab who has been working with coronavirus genetic material and tested positive in June. “It went further than we would ever imagine. My backpack has been continuously positive this entire time.”
The fact that test primers and probe sequences are quite common in human and microbial genetic material may go far in explaining these strange results, as well as the equally strange false positives in lab troubleshooting the first version of CDC’s PCR test in February.
Technical appendix
I asked Dr. Lee the fairly obvious question: “how could it be that these tests could be developed with probes and primers that are common in human DNA and microbes and none of those developers thought to search BLAST for these sequences? Was it just gross negligence or were the test developers somehow not aware that only a 10-base sequence can yield matches and amplification? Is this not a widely accepted view?”
His reply:
The PCR test developers at CDC claim that you need to have a nontarget DNA with two partially matching primer sites at two ends and one partially matching probe site in between on one strand of dsDNA to produce a false positive result. And the 3' end of the primer must not “overhang”. Otherwise, exponential duplication of DNA (PCR) would not take place. So, you may have many partially matching sequences, as you show. But very few, or none, meet all these requirements. So, in theory they can get away with it. Until Sanger sequencing comes in. Demand them to show the sequence in the PCR products.
